Multiple linear regression
We use the barley QTL mapping data to illustrate this function. After deleting unwanted traits and removing irrelevant markers, we are ready to conduct multiple linear regression. (Note: these previous steps are not mandatory, although they simplifies the analysis.)
Under the main menu of Association, click Multiple Linear Regression, and select Conduct Regression. A dependent variable selector will appear on the top of the biplot.
Select the variable that you want to use as the dependent variable, here KW in this example, and click the Regress button. If the number of variables (testers) plus 2 is greater than the number of observations (entries), you will get the following message:

In our case, we had 145 observations, many more than the number of variables. So this was not a problem and multiple regression was conducted as requested. The regression results were printed to the log file, and the biplot was changed to the following:
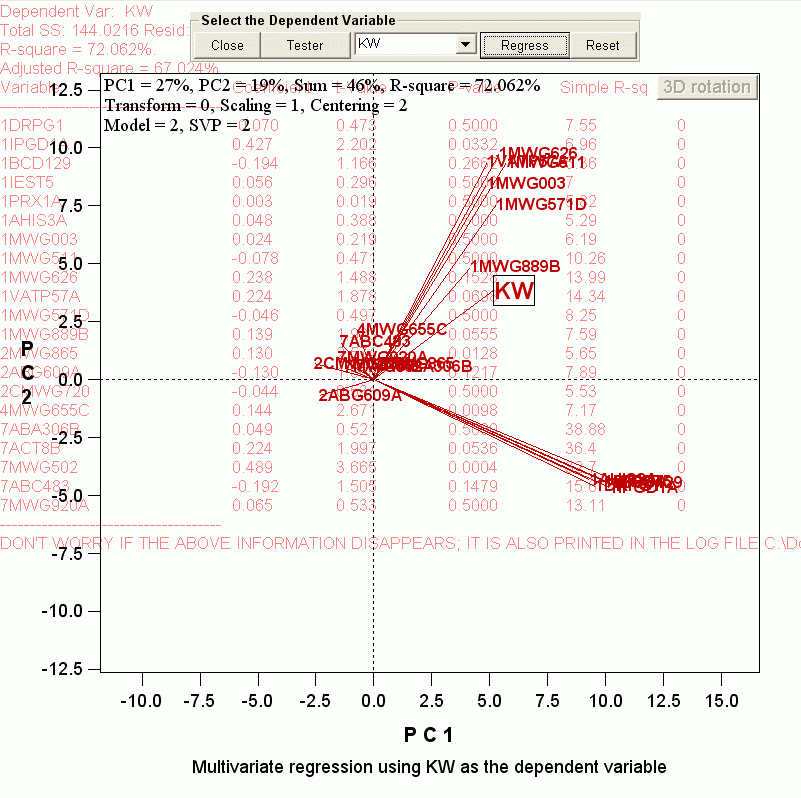
There is much to say about this "biplot" (actually, the above image is not a biplot; it is just a "plot" of the testers, because the genotypes are hidden so that a better view about the "testers" can be achieved).
In Depth:
GGEbiplot has a function to remove markers that are only remotely related to the QTL.